To view this blog on my blog site where you will have access to the other blogs and can sign up for email notifications, please click here.
Let me just say again that the Prostate Cancer Foundation Annual Retreat is overwhelmingly my favorite meeting of the year. The organizers bring in the very best researchers, both clinical and basic science oriented for a two day intense update. Trying to take notes during the sessions leaves you with writer’s cramp and brain ache, but each year brings incredible progress. You can get some idea of the progress for this year from an excellent overview lecture given by Jonathan Simons. Thanks to PCF for getting this up on the web quickly! If you are a patient or family member, please spend 30 minutes viewing Jonathan’s talk and you will definitely not need to ask your physician, “is there anything new?”. (But try not to embarrass him/her with your new insights, either!)
I think the facet of cancer most commonly misunderstood by non-scientists (patients/families) is heterogeneity. While there were several presentations that touched on this, Steven Bova’s talk, reviewing the Nature paper he and his colleagues published this year, probably illustrates the phenomenon best. This international group was able to do whole genome sequencing on cancer metastases (and primary tumors) in 10 patients who had just died from prostate cancer. You can think of this as looking at a fingerprint of different small pieces of tumor found in the prostate, a lymph node in the pelvis, another in the neck area, and maybe two more from some spots you identified on a bone scan. When you look at your fingerprint, there are lines that divide, curve, end, and so forth in patterns that are completely unique to you. Thus each individual tumor metastasis could be identified by the unique set of mutations found at that site and you could look back at the primary tumor to see how many of the mutations and other genetic alterations (deletions, amplifications, etc.) were present in the original tumor or were shared (or not) by another metastasis. What emerges from such investigation is a phenomenal picture of genetic instability starting even in the primary tumor (about which we simply say, “all right, you have a Gleason 3+4 cancer in 3 out of 12 biopsy cores” – an incredible oversimplification of what is really there under the microscope) You can also think of it as tree trunk with multiple branches, then leaves. Here is an illustration from the paper for two patients.
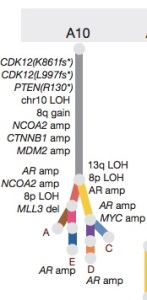 This is the trunk and tree illustration of genetic alterations found in a patient’s metastases. As you can see in the trunk (site of the original tumor) there are already a host of changes, each of which might be related to tumor suppressor genes or oncogenes that drive the tumor. You can appreciate how much more complex this is than simply saying “Gleason pattern 3+4”, and if you look in the same tumor, just a few millimeters away, you might find different alterations. Indeed, in some studies like this, it was the “not dangerous” Gleason 6 pattern tumor from one area of the prostate that spread and led to metastases, not the “more dangerous” Gleason 7 pattern tumor found somewhere else in the patient’s prostate. This gives you some notion of why it may be challenging to send off a bit of tumor for genetic analysis, then make a clinical decision based on just looking at “the worst” area for predicting what to do in terms of treatment. In the figure, you can also appreciate that if you spent millions of dollars finding a drug that worked for the “CTNNB1” amplification, it would only be part of the story. However, it might be a better drug to develop for this patient than one for MYC amplification shown at the lower right, since that target is only present in one branch of the tree. Also note that androgen receptor amplification (AR amp) is found in the “leaves” , illustrating that the tumors in sites D and E have figured out a way to get around hormone treatment by increasing the number of receptors for the tiny amount of testosterone that might not be suppressed by drugs like leuprolide/goserelin/abiraterone or blocked by drugs like bicalutamide/enzalutamide.
This is the trunk and tree illustration of genetic alterations found in a patient’s metastases. As you can see in the trunk (site of the original tumor) there are already a host of changes, each of which might be related to tumor suppressor genes or oncogenes that drive the tumor. You can appreciate how much more complex this is than simply saying “Gleason pattern 3+4”, and if you look in the same tumor, just a few millimeters away, you might find different alterations. Indeed, in some studies like this, it was the “not dangerous” Gleason 6 pattern tumor from one area of the prostate that spread and led to metastases, not the “more dangerous” Gleason 7 pattern tumor found somewhere else in the patient’s prostate. This gives you some notion of why it may be challenging to send off a bit of tumor for genetic analysis, then make a clinical decision based on just looking at “the worst” area for predicting what to do in terms of treatment. In the figure, you can also appreciate that if you spent millions of dollars finding a drug that worked for the “CTNNB1” amplification, it would only be part of the story. However, it might be a better drug to develop for this patient than one for MYC amplification shown at the lower right, since that target is only present in one branch of the tree. Also note that androgen receptor amplification (AR amp) is found in the “leaves” , illustrating that the tumors in sites D and E have figured out a way to get around hormone treatment by increasing the number of receptors for the tiny amount of testosterone that might not be suppressed by drugs like leuprolide/goserelin/abiraterone or blocked by drugs like bicalutamide/enzalutamide.
In this figure, we can see the incredible picture of how the cancer has spread from the prostate to different sites, but also from one site to another, and even from a metastatic site back to the prostate itself. During the conference, there were numerous similar examples of heterogeneity shown. As one example, using the most modern of imaging techniques (such as PET scanning), patients who were initially treated with hormone therapy, and who had “complete remission” as determined by dramatic drops in PSA and disappearance of all metastases on routine bone scans, were in fact shown to have mixed responses with the more sensitive scans. While most of the spots on the sensitive scan went away, a few got worse, and actually new ones appeared elsewhere – all the while, we clinicians are telling patients that “everything looks great” because the regular bone scans and PSA have responded so nicely.
I think all of this leads to the conclusion that no one will ever walk out of a lab with a single new drug that represents “the cure” for all patients with prostate cancer. Instead, we should be able to figure out how to better attack the trunk of the trees for individual patients, rather than just picking off a leaf. For now, the trunk target most tractable relates to the androgen signaling pathway, and the good news is that with the newer drugs available, we seem to be making real progress in turning prostate cancer into more of a chronic disease like diabetes, even if “cure” remains elusive. For many men, the remissions that are available with existing drugs are “good enough” to extend life for many years, sometimes decades. And as I sometimes joke with my patients (in my black humor sort of way… but there is truth here, none-the-less) “If you die of a heart attack at age 90, I’m going to count that as a cure!”





This guy is one of the best in the prostate cancer business. Unfortunately, I think he is retiring.
Sent from my Verizon Wireless 4G LTE smartphone
You are very kind. I am planning to continue to see some patients at the Shaw Cancer Center (Vail) for a few years, and so long as I think I have something to say that might be useful, I’ll continue to blog.
Happy retirement! Glad to see you haven’t stopped learning. Perhaps you already know , but Dave has finished his last study, and beginning pain management through hospice. We cannot thank you enough for all you have done!!!
Only wish there was more to do. Best wishes.
I will miss you and your quick responses to my many questions. It will be great if you continue to blog.
This one was extremely helpful to understand why my “Gleason 6” tumor metastasized and has not been predictable.
Thanks for all the help you gave over the years.
Given this heterogeneity of tumors, especially in prostate cancer, I can’t help but wonder if all the time and money we’ve pumped into research on the genetic basis for cancer over the last several decades was well spent. It’s fascinating stuff, thanks for keeping up the blog.
Pingback: Olaparib for resistant prostate cancer | prost8blog
Pingback: 3 Articles and a forth | prost8blog
Pingback: The Hits Just Keep on Coming | prost8blog
Pingback: Why can’t we cure this??? | prost8blog
Pingback: Epigenetics | prost8blog